Recurrent Breaking Genes in Neural Stem and Progenitor Cells
Date:
Wednesday, February 21, 2018 10:00 - 11:00
Speaker:
Peggy Wei (Boston Children's Hospital)
Location:
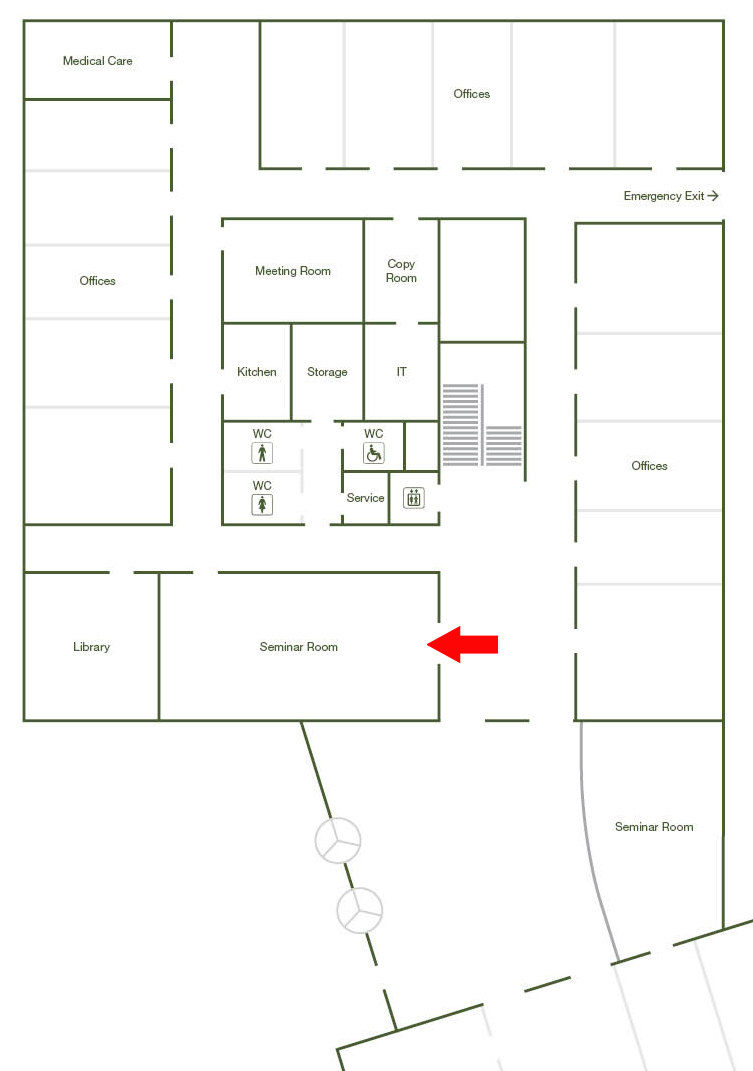
Big Seminar room Ground floor / Office Bldg West (I21.EG.101)
Series:
Life Sciences Seminar
Host:
Simon Hippenmeyer
Contact:
DEL REAL LAVERGNE Pedro
Repair of DNA double strand breaks (DSBs) by the classical non-homologous end joining (C-NHEJ) pathway is required for brain development. In mice, neural stem and progenitor cell (NSPC)-specific inactivation of C-NHEJ ligation complex in a p53-deficient background leads to medulloblastomas, the most common and lethal pediatric brain cancer. To identify the source of DSBs, we developed high throughput, genome-wide, translocation sequencing (HTGTS) to map DSBs in base-pair resolution, and found 113 recurrent DSB clusters (RDCs) that occur in transcribed genes in murine NSPCs. The majority of RDC-genes are involved in synaptic function and have genetic implications in brain development, neuropsychiatric diseases, and cancer. RDC-genes are generally very long with short exons, and they often lie within topological associated domains (TADs). Frequent joining of intronic DSBs coupling with exon loss is found in RDC-genes. Alternative spliced-like RDC-gene transcripts that encode such hard-wired gene templates were also identified. We propose a new mechanism which could explain further neuronal transcript diversification in development, and its dysregulation to cause brain disease.