Exploiting data structure: from gene expression to evolution
Date:
Tuesday, March 6, 2018 09:00 - 10:00
Speaker:
Maria Chikina (U of Pittsburgh)
Location:
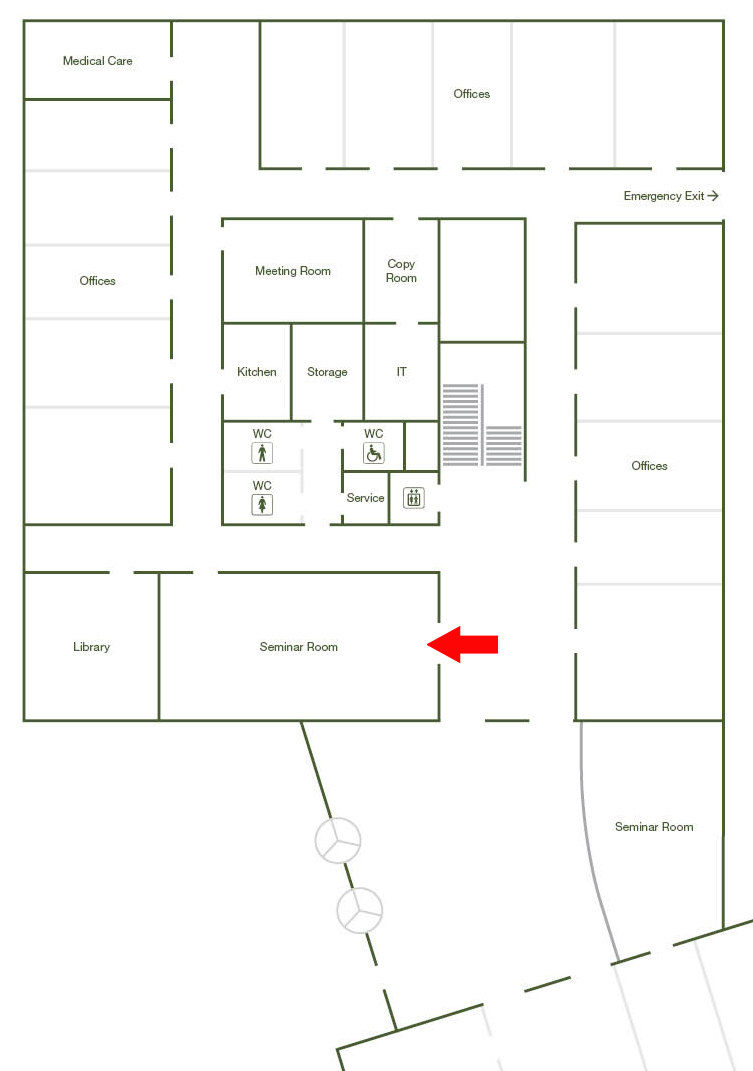
Big Seminar room Ground floor / Office Bldg West (I21.EG.101)
Series:
Life Sciences Seminar
Host:
Nick Barton
Contact:
DEL REAL LAVERGNE Pedro
Genome scale molecular datasets are often highly structured, with many correlated observations. This general phenomenon can be related to the underlying data generating process. In gene expression assays, groups of gene are co-regulated through shared transcription factors and signaling pathways. Likewise, for evolutionary data, correlated substitution rates arise due to shared evolutionary pressure. In the first half of the talk we will present a new constrained matrix decomposition approach that directly aligns a lower dimension representation with known biological pathways. Our method provides state-of-the-art accuracy in reconstructing known upstream variables through a biologically interpretabile decomposition. In the second half we will discuss a computational approach for modeling the relationship between phenotypes and evolutionary rates and present new results on evolutionary correlates of life-history traits.