Discovering molecular nuts and bolts in the nervous system
Date:
Thursday, March 1, 2018 10:00 - 11:00
Speaker:
Mario de Bono (MRC Lab of Molecular Biology)
Location:
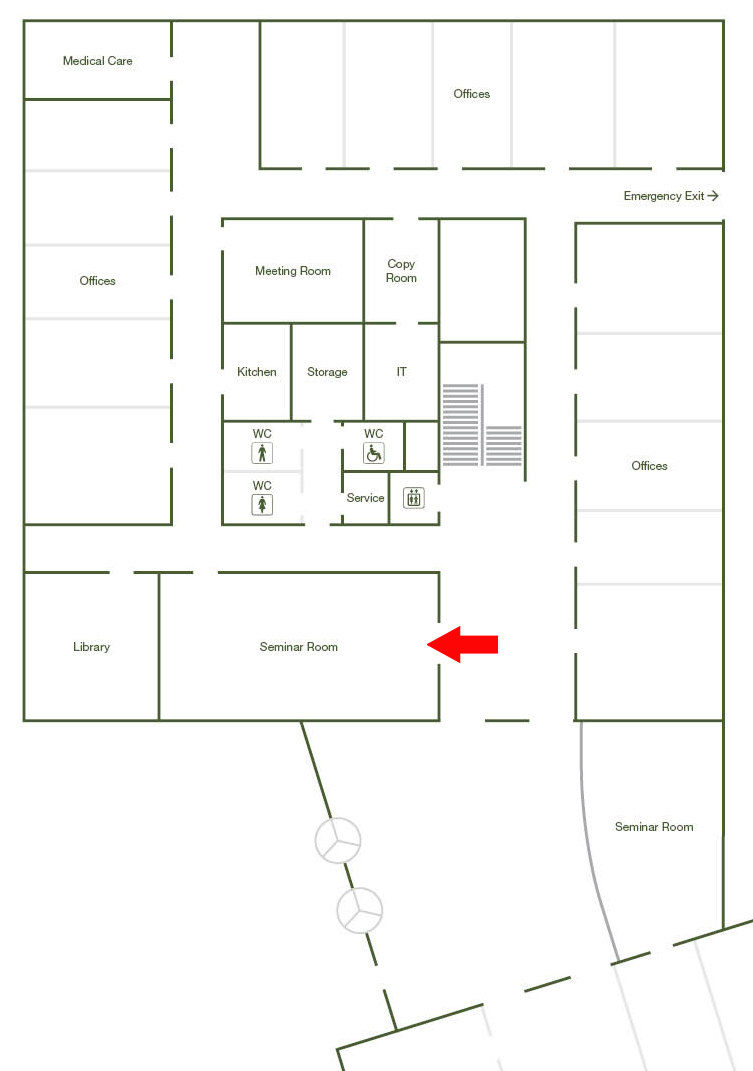
Big Seminar room Ground floor / Office Bldg West (I21.EG.101)
Series:
Life Sciences Seminar
Host:
Peter Jonas
Contact:
DEL REAL LAVERGNE Pedro
We aim to discover molecular mechanisms that underpin neuron and circuit properties. As entry points we identify mutants in the nematode C. elegans unable to adopt particular behavioral states. Forward genetics is a potent tool for discovering biological mechanism. However, linking phenotypes to mutations requires laborious mapping. We circumvent this by showing that bioinformatics alone can predict phenotype-causing mutations in large collections of sequenced mutants. From 251 mutants defective in C. elegans aggregation behavior, we identify (thus far) 43 genes promoting this behavior. The proteins they encode define several molecular machines. All act in the same defined circuit. I will describe some of our discoveries. Unexpectedly, we find that interleukin 17, a pro-inflammatory cytokine, can act on neurons to change circuit gain. We identify complexes in the endoplasmic reticulum that appear to control biogenesis of ion channels and GPCRs. These complexes are conserved, and we are using biochemistry to understand their function in mammalian cells.