Evolutionary genomics in centromeric and pericentromeric regions of Drosophila and humans
Date:
Monday, January 14, 2019 11:00 - 12:00
Speaker:
Chuck Langley (University of California, Davis)
Location:
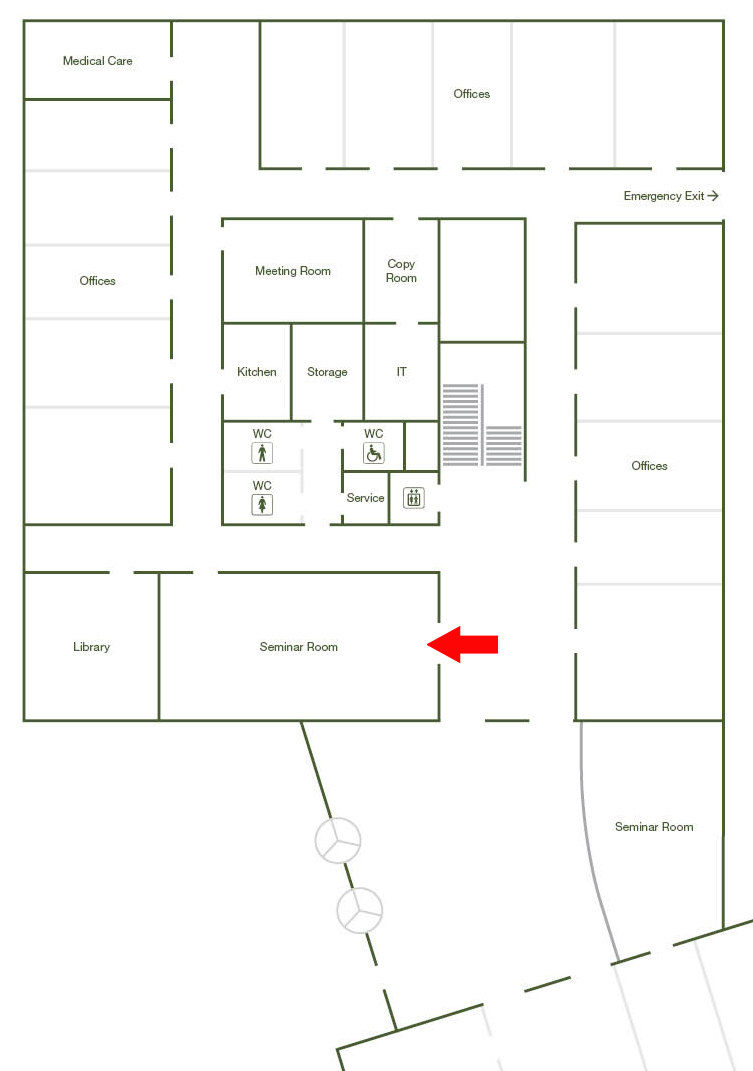
Big Seminar room Ground floor / Office Bldg West (I21.EG.101)
Series:
Life Sciences Seminar
Host:
Nick Barton; Beatriz Vicoso
Contact:
HACKER Elisabeth
Centromeres and their surrounding pericentric heterochromatic regions remain enigmatic and poorly understood despite critical roles in eukaryotic inheritance. Their vast size, highly repetitive structure, paucity of coding genes and low recombination rates have impeded genetic and genomic advancement. Potentially large selective impacts of recurrent meiotic drive in female meiosis have been proffered as the causes of evolutionarily rapid genomic 'turnover' centromere associated satellite DNAs. The epigenetic inheritance of the functional centromere and the growing catalog of the genes/proteins and pathways essential to centromeres function all motivate a full accounting of genomic diversity in centromeric regions. I shall report on identification of cenhaps, large scale haplotypic variation in both humans and Drosophila that spans the complete centromerics regions of metacentic chromosomes, including the unassembled gaps in the reference genomes that contain Mbp of highly repeated sequences (170 bp alpha-satellites in humans and smaller satellites in flies). The inferred dynamics behind the apparent descent of cenhaps are surprisingly rich and complex, including archaic lineages. Variation among cenhaps in satellite DNA content is common, opening a direct avenue to test hypotheses about the mechanistic consequence of heterozygosity for large differences in 'centromere strength' in mitosis and meiosis, as well as to more precise modeling of their evolutionary dynamics.